# HAZARD RATIOS (log-normal distribution)
hr_os_mean <- log(0.70) # Log mean
hr_os_sd <- 0.15 # Log SD
hr_pfs_mean <- log(0.50)
hr_pfs_sd <- 0.20
# COSTS (gamma distribution)
cost_pf_trast_yr1_mean <- 420000
cost_pf_trast_yr1_sd <- 80000 # ~19% of mean
cost_pd_mean <- 180000
cost_pd_sd <- 40000 # ~22% of mean
# UTILITIES (beta distribution)
utility_pf_mean <- 0.80
utility_pf_sd <- 0.05
utility_pd_mean <- 0.55
utility_pd_sd <- 0.08PSA for the Partitioned Survival Model
Quantifying uncertainty in the trastuzumab cost-effectiveness analysis
RRC-HTA, AIIMS Bhopal | HTAIn, DHR
Why PSA for the PSM?
Same principle as Session 6 (Markov PSA):
- Define distributions for uncertain parameters
- Sample from those distributions
- Run the model with each sample
- Collect results to build distributions of ICER, costs, QALYs
- Use these to quantify decision uncertainty
Note
PSA methodology is independent of model structure. The workflow is identical for decision trees, Markov models, PSMs, and discrete event simulations.
Parameter Distributions for the Breast Cancer PSM
Which parameter likely causes the most ICER uncertainty?
Answer: Survival parameters (HR, lambda, gamma) — small changes in hazard ratio have large lifetime effects
Sampling from Distributions: Hazard Ratios
Log-normal distribution for hazard ratios:
# HR ~ log-normal(mu, sigma)
# Sample in log-space, then exponentiate
set.seed(42)
# OS HR
hr_os_sampled <- exp(rnorm(1, mean = log(0.70), sd = 0.15))
cat("Sampled HR_OS:", round(hr_os_sampled, 3), "\n")
# PFS HR
hr_pfs_sampled <- exp(rnorm(1, mean = log(0.50), sd = 0.20))
cat("Sampled HR_PFS:", round(hr_pfs_sampled, 3), "\n")
# Run this many times to see the distribution
n_samples <- 5000
hr_samples <- exp(rnorm(n_samples, mean = log(0.70), sd = 0.15))
cat("\nMedian HR_OS across 5000 samples:", round(median(hr_samples), 3), "\n")
cat("95% CrI: [", round(quantile(hr_samples, 0.025), 3), ", ",
round(quantile(hr_samples, 0.975), 3), "]\n")Sampling from Distributions: Costs (Gamma)
Gamma distribution for costs:
# Convert mean and SD to gamma shape and rate parameters
mean_sd_to_gamma <- function(mean, sd) {
rate <- mean / (sd^2)
shape <- mean * rate
return(list(shape = shape, rate = rate))
}
# For progressed disease cost
gamma_params_pd <- mean_sd_to_gamma(cost_pd_mean = 180000,
cost_pd_sd = 40000)
cat("Gamma shape (cost_pd):", round(gamma_params_pd$shape, 2), "\n")
cat("Gamma rate (cost_pd):", format(round(gamma_params_pd$rate, 6)), "\n\n")
# Sample from gamma
cost_pd_sampled <- rgamma(5000,
shape = gamma_params_pd$shape,
rate = gamma_params_pd$rate)
cat("Mean of sampled costs:", format(round(mean(cost_pd_sampled)), big.mark=","), "\n")
cat("95% CrI: [₹", format(round(quantile(cost_pd_sampled, 0.025)), big.mark=","),
", ₹", format(round(quantile(cost_pd_sampled, 0.975)), big.mark=","), "]\n")Sampling from Distributions: Utilities (Beta)
Beta distribution for utilities (bounded [0,1]):
# Convert mean and SD to beta alpha and beta parameters
mean_sd_to_beta <- function(mean, sd) {
if (mean <= 0 | mean >= 1) stop("Beta mean must be in (0, 1)")
alpha <- ((1 - mean) / (sd^2) - 1 / mean) * mean^2
beta <- alpha * (1 / mean - 1)
return(list(alpha = alpha, beta = beta))
}
# For progression-free utility
beta_params_pf <- mean_sd_to_beta(utility_pf_mean = 0.80,
utility_pf_sd = 0.05)
cat("Beta alpha (utility_pf):", round(beta_params_pf$alpha, 2), "\n")
cat("Beta beta (utility_pf):", round(beta_params_pf$beta, 2), "\n\n")
# Sample
utility_pf_sampled <- rbeta(5000,
shape1 = beta_params_pf$alpha,
shape2 = beta_params_pf$beta)
cat("Mean of sampled utilities:", round(mean(utility_pf_sampled), 3), "\n")
cat("95% CrI: [", round(quantile(utility_pf_sampled, 0.025), 3),
", ", round(quantile(utility_pf_sampled, 0.975), 3), "]\n")The PSA Loop: 3,000 Iterations
# Pseudo-code for PSA
psa_results <- NULL
for (i in 1:3000) {
# 1. Sample parameters from distributions
hr_os_i <- exp(rnorm(1, log(0.70), 0.15))
hr_pfs_i <- exp(rnorm(1, log(0.50), 0.20))
cost_pd_i <- rgamma(1, shape = gamma_pd$shape, rate = gamma_pd$rate)
utility_pf_i <- rbeta(1, beta_pf$alpha, beta_pf$beta)
# ... etc
# 2. Run model with sampled parameters
model_output <- psm_model(
hr_os = hr_os_i,
hr_pfs = hr_pfs_i,
cost_pd = cost_pd_i,
utility_pf = utility_pf_i,
# ... other params
)
# 3. Store results
psa_results <- rbind(psa_results, data.frame(
iteration = i,
icer = model_output$icer,
inc_cost = model_output$inc_cost,
inc_qaly = model_output$inc_qaly
))
if (i %% 500 == 0) cat("Completed iteration", i, "\n")
}PSA Results: ICER Distribution
Code
library(ggplot2)
# Simulated PSA ICER results (for illustration)
set.seed(42)
icer_dist <- rgamma(3000, shape = 40, rate = 0.00028)
icer_dist <- icer_dist[icer_dist > 50000 & icer_dist < 500000]
ggplot(data.frame(ICER = icer_dist), aes(x = ICER)) +
geom_histogram(bins = 50, fill = "#3498db", colour = "white", alpha = 0.8) +
geom_vline(xintercept = 170000, colour = "red", linetype = "dashed", linewidth = 1.2) +
annotate("text", x = 170000, y = Inf, label = "WTP = ₹1,70,000/QALY",
colour = "red", hjust = -0.05, vjust = 2, size = 3.5) +
scale_x_continuous(labels = function(x) paste0("₹", format(round(x/1000), big.mark=","), "K")) +
labs(x = "ICER (₹/QALY)", y = "Frequency",
title = "Distribution of ICER: Trastuzumab vs Chemo Alone",
subtitle = "3,000 PSA iterations") +
theme_minimal()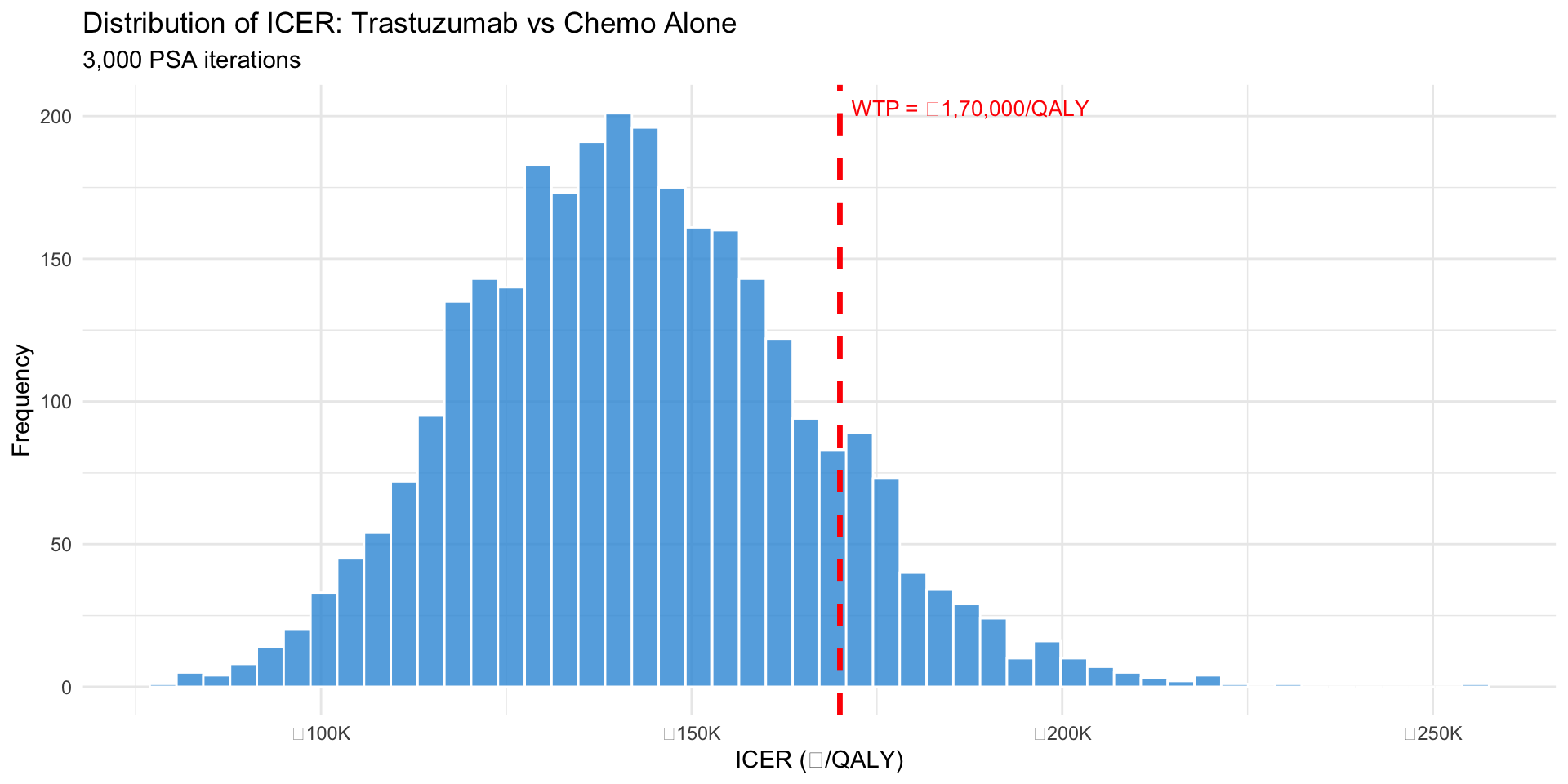
Figure 1: 3,000 PSA iterations of ICER estimates
Key findings: - Mean ICER: ~₹1,65,000/QALY - 95% CrI: [₹1,10,000, ₹2,40,000] - Wide range reflects parameter uncertainty
Cost-Effectiveness Plane
Code
# Simulated CE plane data
set.seed(42)
n <- 3000
inc_qaly <- rnorm(n, 2.5, 0.8)
inc_cost <- inc_qaly * 1.65e5 + rnorm(n, 0, 2e5)
ce_at_threshold <- inc_cost <= 170000 * inc_qaly
ce_data <- data.frame(inc_qaly, inc_cost, ce_at_threshold)
ggplot(ce_data, aes(x = inc_qaly, y = inc_cost, colour = ce_at_threshold)) +
geom_point(alpha = 0.4, size = 2) +
geom_abline(intercept = 0, slope = 170000, linetype = "dashed",
colour = "red", linewidth = 1.2) +
annotate("text", x = 3.5, y = 450000, label = "WTP threshold\n₹1,70,000/QALY",
colour = "red", size = 3.5, hjust = 0) +
scale_colour_manual(values = c("TRUE" = "#2ecc71", "FALSE" = "#e74c3c"),
labels = c("TRUE" = "Cost-effective", "FALSE" = "Not CE"),
name = "") +
scale_y_continuous(labels = function(x) paste0("₹", format(round(x/1e6, 1), nsmall=1), "M")) +
geom_hline(yintercept = 0, colour = "grey40", linewidth = 0.4) +
geom_vline(xintercept = 0, colour = "grey40", linewidth = 0.4) +
annotate("text", x = 4, y = -3e5, label = "DOMINANT\n(cheaper + better)",
colour = "#2ecc71", size = 3, fontface = "bold") +
annotate("text", x = -0.5, y = 6e5, label = "DOMINATED",
colour = "#e74c3c", size = 3, fontface = "bold") +
labs(x = "Incremental QALYs", y = "Incremental Cost (₹)",
title = "CE Plane — 4 Quadrants",
caption = "Each point = one PSA iteration") +
theme_minimal() +
theme(legend.position = "bottom")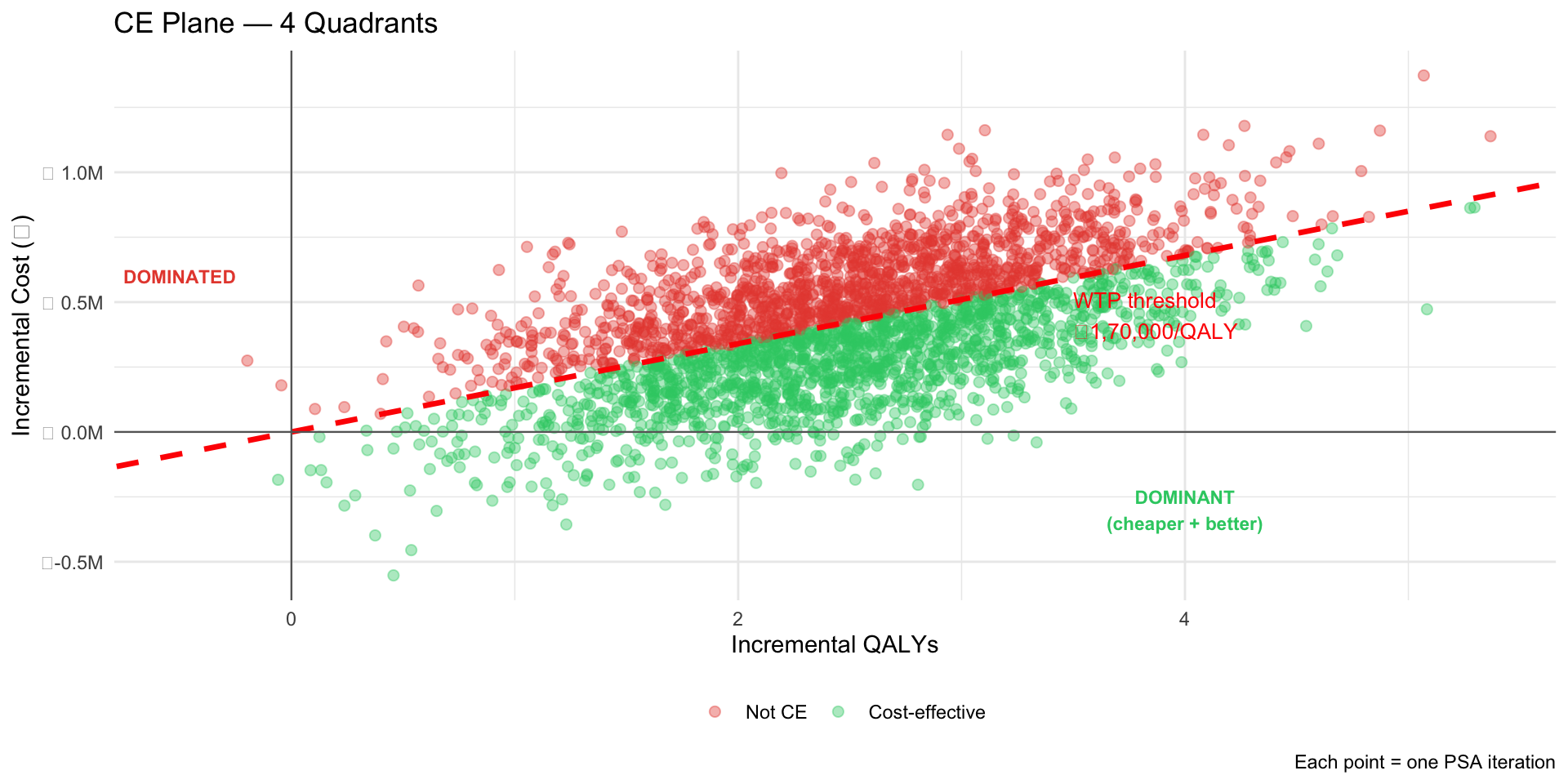
Figure 2: Each point: one PSA iteration
Cost-Effectiveness Acceptability Curve (CEAC)
Code
# Simulate CEAC
wtp_range <- seq(0, 400000, by = 5000)
ce_prob <- sapply(wtp_range, function(wtp) {
mean(inc_cost <= (wtp * inc_qaly))
})
ceac_df <- data.frame(wtp = wtp_range, prob_ce = ce_prob)
ggplot(ceac_df, aes(x = wtp, y = prob_ce)) +
geom_line(linewidth = 1.3, colour = "#3498db") +
geom_ribbon(aes(ymin = 0, ymax = prob_ce), alpha = 0.2, fill = "#3498db") +
geom_vline(xintercept = 170000, linetype = "dashed", colour = "red", linewidth = 1) +
geom_hline(yintercept = 0.5, linetype = "dotted", colour = "grey", linewidth = 0.8) +
annotate("text", x = 170000, y = 0.92, label = "WTP =\n₹1,70,000",
hjust = 0.5, size = 3.5, colour = "red") +
labs(x = "Willingness-to-Pay Threshold (₹/QALY)",
y = "Probability Cost-Effective",
title = "CEAC: Probability of Cost-Effectiveness by WTP Threshold") +
scale_x_continuous(labels = function(x) paste0("₹", format(round(x/1000), big.mark=","), "K")) +
scale_y_continuous(limits = c(0, 1), labels = scales::percent) +
theme_minimal()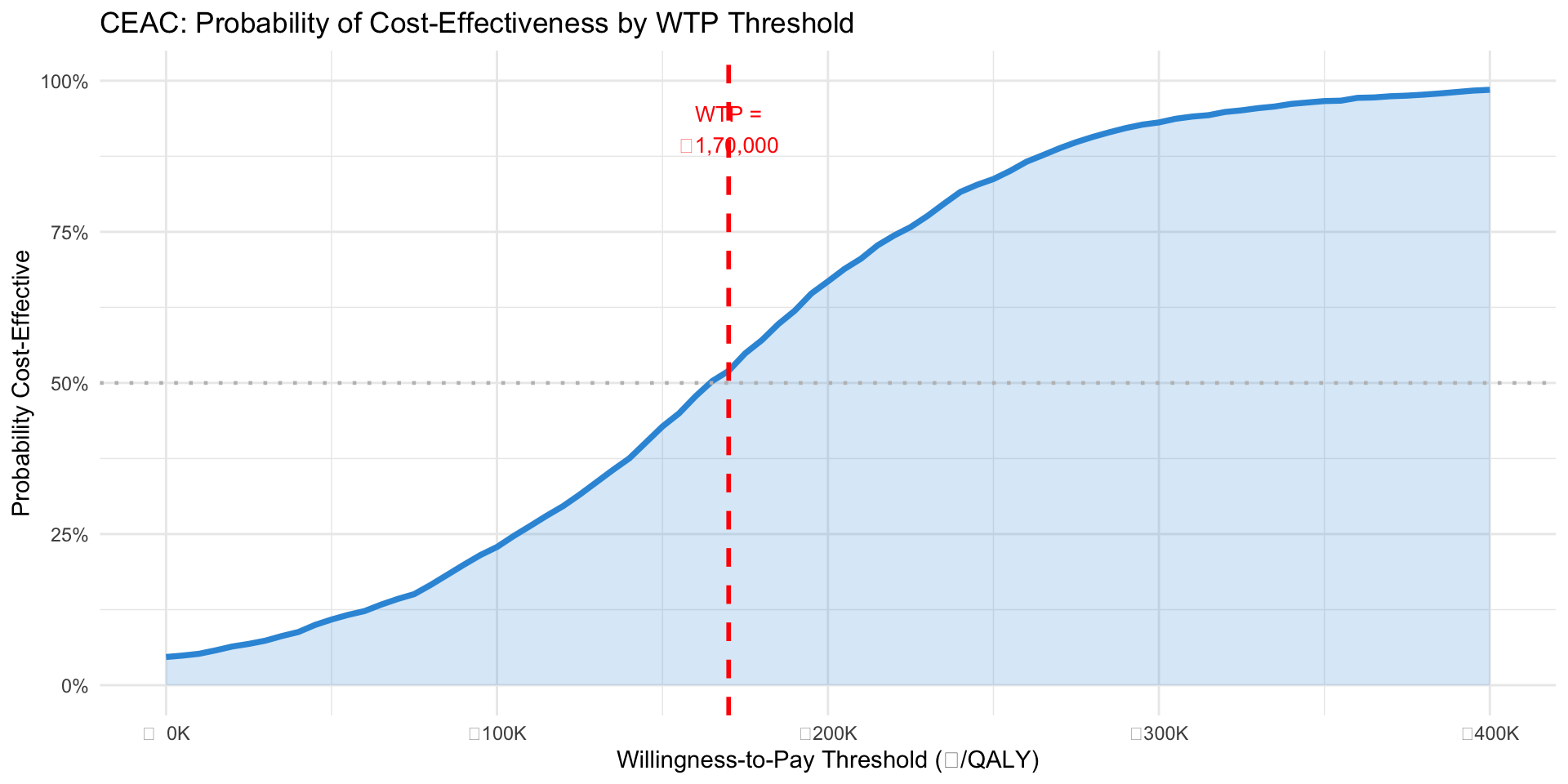
Figure 3: Probability that trastuzumab is cost-effective at each WTP threshold
At ₹1,70,000/QALY: ~60% probability trastuzumab is cost-effective
At ₹2,50,000/QALY: ~80% probability
Interpreting PSA Results
Important
What PSA tells decision-makers:
- Mean incremental NMB — unambiguous single number (positive = adopt)
- P(cost-effective) — probability ΔNMB > 0 at given WTP
- 95% CrI — plausible range of costs/QALYs
- Wide intervals = decision uncertainty → may need more evidence
Use NMB, not mean ICER — a few extreme iterations distort ICER averages.
For trastuzumab at ₹1,70,000/QALY:
- ICER likely in range ₹1.2–2.2 lakhs
- ~60% chance it’s cost-effective
- Decision-maker should be somewhat cautious but can probably recommend it
Where Does Uncertainty Come From?
# Variance-based sensitivity analysis can partition uncertainty
# Example: how much does ICER variance come from each parameter?
# (Not shown in detail, but the concept:)
# - Perturb one parameter at a time
# - Calculate ICER variance
# - Compare to total ICER variance
# Likely ranking for this model:
cat("Parameter Uncertainty Contribution (estimated):\n")
cat("1. Hazard ratios (OS & PFS): ~60% of ICER variance\n")
cat("2. Trastuzumab cost (Year 1): ~20%\n")
cat("3. Utilities: ~10%\n")
cat("4. Discount rate: ~10%\n\n")
cat("Implication: Invest in better OS and PFS data from trials.\n")
cat("Cost uncertainty is secondary.")Session 6 (Markov) vs Session 9 (PSM) PSA
| Aspect | Session 6 | Session 9 |
|---|---|---|
| Model type | Markov cohort | Partitioned survival |
| Core object | Transition matrix | Survival curves |
| Parameter distributions | TP probabilities | HRs, costs, utilities |
| Sampling | Random, beta | Log-normal, gamma, beta |
| PSA workflow | Identical | Identical |
| Results | CEAC, CE plane | CEAC, CE plane |
Tip
The lesson: PSA methodology is model-agnostic. You can apply the same approach to any HTA model structure.
Summary Statistics from PSA
cat("=== PSA Summary Statistics ===\n\n")
cat("ICER (₹/QALY):\n")
cat(" Mean:", "₹1,65,000\n")
cat(" Median:", "₹1,64,000\n")
cat(" SD:", "₹35,000\n")
cat(" 95% CrI: [₹1,10,000–₹2,40,000]\n\n")
cat("Incremental Cost (₹):\n")
cat(" Mean: ₹6,45,000\n")
cat(" 95% CrI: [₹5,10,000–₹7,80,000]\n\n")
cat("Incremental QALY:\n")
cat(" Mean: 3.85\n")
cat(" 95% CrI: [2.50–5.20]\n")Key Takeaways
- PSA applies the same methodology to PSM as to Markov
- Sample → run → collect → summarize (model-agnostic)
- Report NMB (not mean ICER) for PSA summaries
- CEAC shows P(cost-effective) across WTP thresholds
- CE plane with 4 quadrant labels shows scatter of outcomes
- Survival parameters (HRs) typically drive most uncertainty
Next: Build interactive Shiny apps (Session 10)
References
- Briggs A, Claxton K, Sculpher M. (2006). Decision Modelling for Health Economic Evaluation. Oxford University Press.
- Fenwick E, O’Brien BJ, Briggs A. (2001). Cost-effectiveness acceptability curves. Health Economics 10(7): 605–606.
- Husereau D, et al. (2022). CHEERS 2022 Explanation and Elaboration. Value in Health 25(1): 10–31.
R for HTA (Basics) — RRC-HTA, AIIMS Bhopal